Introduction
MBioSEQ Ridom Typer implements CheckM2 (citation), a machine learning–based technique to evaluate microbial genome assemblies by identifying intra-species contamination and predicting genome completeness. CheckM2 delivers highly accurate quality predictions for medium- and low-quality genomes, including those from poorly represented lineages. It can even produce reliable results for phyla with only a single genomic representative.
CheckM2 uses two machine learning models to predict genome completeness score. A 'general' gradient boosting model is designed for novel or distantly related organisms (e.g., new orders, classes, or phyla) and generalizes well to poorly represented lineages. A 'specific' neural network model provides higher accuracy for genomes closely related to the reference set (e.g., known species, genera, or families), particularly when genomes are less complete. This ensures better accuracy across lineages.
Please note: Especially with older Illumina machines there is frequently some low contamination due to inter-run valve and/or flexible tube carry-over. Notable the SKESA assembler handles contaminations of up to 10% very well.
Task Entry Overview
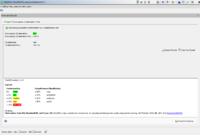
The task entry overview shows the CheckM2 results for the sample.
The Intra-species Contamination is color-coded according to the quantitative value of contamination percentage using the following thresholds:
- Green: Intra-species Contamination Perc. ≤5%
- Yellow: Intra-species Contamination Perc. >5% and ≤15%
- Red: Intra-species Contamination Perc. >15%
Result Fields
The task entry stores the following result fields for each sample:
| Field | Description |
|---|---|
| Intra-species Contamination | A classification based on the value for Intra-species Contamination Percentage. For QC this field is highlighted green if the contamination is below 5%, yellow if it is between 5% and 15%, and red if it is above 15%. |
| Intra-species Contamination Perc. | Estimated percentage of contamination (redundant or foreign sequences) in the assembly consensus. |
| Completeness | A classification based on the value for Completeness Percentage |
| Completeness Perc. | Estimated percentage of the genome recovered, predicted by CheckM2's machine learning model. |
The controlled vocabulary that is shown as legend in the results for classification of draft genome quality based on estimated genome completeness and contamination is the one used by the developers in their first publication (Parks et al. Genome Res. 2015, 25: 1043-55 [PubMed 25977477]).
The Result tab of the Sample Overview shows the Intra-species Contamination and the Completeness fields. Those two fields are also written to the Procedure Statistics. When a comparison table is created for a project that contains the CheckM2 Intra-species Contamination Check Task Template the two fields are automatically added to the comparison table. If the task template is explicitly selected when creating a comparison table, all result fields are added to the table.
Run times
| Species | NCBI ID | Run time |
|---|---|---|
| Escherichia coli | NC_000913.3 | 1m 14sec |
| Listeria monocytogenes | NC_003210.1 | 58 sec |
| Staphylococcus aureus | NZ_CP007455.1 | 37 sec |
FOR RESEARCH USE ONLY. NOT FOR USE IN CLINICAL DIAGNOSTIC PROCEDURES.